Flux upper limits with NITRATES#
# ---- User inputs ----
work_dir = '/Users/sjs8171/Desktop/acme_tutorials/nitrates/785996578_c0'
temporal_bin_s = 16
sigma_ul = 5.0
nside_out = 32
output_fits = f'{work_dir}/ul_skymap.fits'
# Paths to UL response files (.rsp)
resp_dir_override = '/Users/sjs8171/Desktop/NITRATES/tests/nitrates_resp_dir/rsps4limits2/'
#resp_dir_override_2 = '/Users/sjs8171/Desktop/NITRATES/tests/nitrates_resp_dir/rsps4limits2/'
# spectrum options: 'band' or 'comp'
spectrum_model = 'band'
# For Band
band_alpha = -1.0
band_beta = -2.3
band_epeak_keV = 1000
# For Comptonized
comp_alpha = -0.62
comp_epeak_keV = 185.0
# flux integration range (keV)
flux_elo_keV = 15.0
flux_ehi_keV = 350.0
import os
import numpy as np
from astropy.io import fits
from astropy.table import Table
import healpy as hp
from scipy.interpolate import LinearNDInterpolator, NearestNDInterpolator
from nitrates.lib.event2dpi_funcs import mask_detxy
from nitrates.lib.sqlite_funcs import get_conn
from nitrates.lib.dbread_funcs import get_info_tab
from nitrates.lib.coord_conv_funcs import convert_theta_phi2radec
from nitrates.analysis_seeds.do_full_rates import Linear_Rates
from nitrates.lib.calc_BAT_ul import get_resp4ul_tab, rate2band_eflux, rate2comp_eflux
from nitrates.config import resp_dname
def build_interpolator(ras, decs, vals):
pts = np.column_stack((ras, decs))
lin = LinearNDInterpolator(pts, vals, fill_value=np.nan)
nn = NearestNDInterpolator(pts, vals)
def f(ra, dec):
v = lin(ra, dec)
if np.isnan(v):
return float(nn(ra, dec))
return float(v)
return f
def compute_rate_upper_limit(bkg_obj, trigger_time, dur, sigma=5.0, dtmin=-20.0, dtmax=20.0):
tstep = dur / 4.0
tbins0 = np.arange(dtmin, dtmax, tstep) + trigger_time
tbins1 = tbins0 + dur
sig2_bkg_vals = []
for t0, t1 in zip(tbins0, tbins1):
d = t1 - t0
tmid = 0.5 * (t0 + t1)
bkg_rate, bkg_rate_err = bkg_obj.get_rate(tmid)
sig2_bkg = (bkg_rate_err * d) ** 2 + (bkg_rate * d)
sig2_bkg_vals.append(sig2_bkg)
rate_std = np.sqrt(np.max(sig2_bkg_vals)) / dur
return sigma * rate_std
Steps behind the flux upper limit (UL)#
This notebook follows the same logic used in UL.py.
Below, each step is linked to the function(s) that perform it.
Background fluctuation for a time bin \(\Delta t\)
For each trial window around trigger time, define\[ \sigma^2_{\mathrm{bkg}} = (\sigma_r\,\Delta t)^2 + r_{\mathrm{bkg}}\,\Delta t \]where \(r_{\mathrm{bkg}}\) and \(\sigma_r\) come from the background fit. The term \(r_{\mathrm{bkg}}\Delta t\) is the variance due to Poissonian fluctuation, while \((\sigma_r\,\Delta t)^2\) is the uncertainty coming from background fit. Hence, a background with larger fluctuations implies less stringent upper limits
Functions used:
Linear_Rates(...).do_fits()builds the background model in time.bkg_obj.get_rate(tmid)returns \(r_{\mathrm{bkg}}\) and \(\sigma_r\) at each trial midpoint.compute_rate_upper_limit(...)computes \(\sigma^2_{\mathrm{bkg}}\) over all windows.
Rate upper limit at significance \(N_\sigma\)
Using the worst-case variance across scanned windows:\[ \sigma_{\mathrm{rate}} = \frac{\sqrt{\max(\sigma^2_{\mathrm{bkg}})}}{\Delta t}, \qquad r_{\mathrm{UL}} = N_\sigma\,\sigma_{\mathrm{rate}} \]In this notebook \(N_\sigma = 5\) by default.
Functions used:
compute_rate_upper_limit(...)returns \(r_{\mathrm{UL}}\).
Convert count-rate UL to energy-flux UL (assumed spectrum)
For each sky direction (each DRM):load the DRM,
scale by active-detector fraction,
assume spectral model (Band or Comptonized),
solve normalization that reproduces \(r_{\mathrm{UL}}\),
integrate in the chosen energy range to get
\[ F_{\mathrm{UL}}\;[\mathrm{erg\,cm^{-2}\,s^{-1}}]. \]Functions used:
get_resp_tab_local(...)loads the response file for each \((\theta,\phi)\).rate2band_eflux(...)converts rate UL to flux UL for a Band spectrum.rate2comp_eflux(...)converts rate UL to flux UL for a Comptonized spectrum.
Sky map construction
Directional values \(F_{\mathrm{UL}}(\alpha,\delta)\) are interpolated over the sphere and sampled on a HEALPix grid, then written to FITS.Functions used:
build_interpolator(ras, decs, vals)builds the sky interpolation function.hp.pix2ang(...)maps HEALPix pixel index to sky coordinates.hp.write_map(...)writes the UL sky map in FITS format.
Read UL at a given sky position
Once the FITS map is written, the UL at any \((\mathrm{RA},\mathrm{Dec})\) is obtained by selecting the corresponding HEALPix pixel.Functions used:
ul_at_radec(...)wrapper for point queries.hp.ang2pix(...)converts \((\mathrm{RA},\mathrm{Dec})\) to pixel index.hp.read_map(...)loads the UL FITS map.
Data loading and response-grid setup#
This section prepares all inputs needed by the UL computation.
Load trigger metadata and event products
Opens
results.dband reads trigger MET.Loads event data, GTIs, detector mask, and attitude.
Functions used:
get_conn(...),get_info_tab(...)fits.open(...),Table.read(...)
Build event selection and background model
Applies quality/energy/time cuts to events.
Fits the background model used later for rate UL.
Functions used:
mask_detxy(...)for detector-quality maskingLinear_Rates(...)andbkg_obj.do_fits()for background fitting
Discover response files (.rsp)
Resolves one or more response directories.
Collects all
.rspfiles and removes duplicated filenames.
Functions used:
os.path.isdir(...),os.listdir(...)
Build sky-direction arrays from response filenames
Extracts \((\theta,\phi)\) from each filename.
Converts to sky coordinates \((\mathrm{RA},\mathrm{Dec})\) for interpolation.
Functions used:
convert_theta_phi2radec(...)
Define local response loader per direction
get_resp_tab_local(theta, phi, rsp_dir)opens the exact DRM for each direction.
Functions used:
Table.read(...)
# ---- Load prepared NITRATES products ----
import nitrates.lib.calc_BAT_ul as calc_bat_ul
conn = get_conn(os.path.join(work_dir, 'results.db'))
info_tab = get_info_tab(conn)
trigger_time = info_tab['trigtimeMET'][0]
evfname = os.path.join(work_dir, 'filter_evdata.fits')
ev_data = fits.open(evfname)[1].data
GTI_PNT = Table.read(evfname, hdu='GTI_POINTING')
dmask = fits.open(os.path.join(work_dir, 'detmask.fits'))[0].data
attfile = fits.open(os.path.join(work_dir, 'attitude.fits'))[1].data
att_q = attfile['QPARAM'][np.argmin(np.abs(attfile['TIME'] - trigger_time))]
ndets_active = np.sum(dmask == 0)
Ndet_ratio = ndets_active / 32768.0
mask_vals = mask_detxy(dmask, ev_data)
bl_ev = (ev_data['EVENT_FLAGS'] < 1) & (ev_data['ENERGY'] >= 15.0) & (ev_data['ENERGY'] <= 350.0) & (mask_vals == 0.0) & (ev_data['TIME'] >= trigger_time - 1e3) & (ev_data['TIME'] <= trigger_time + 1e3)
ev_data0 = ev_data[bl_ev]
tmin = GTI_PNT['START'][0]
tmax = GTI_PNT['STOP'][-1]
bkg_obj = Linear_Rates(ev_data0, tmin, tmax, trigger_time, GTI_PNT, sig_clip=4.0, poly_trng=20)
bkg_obj.do_fits()
# response grid directions (supports one or two folders)
resp_dir_candidates = []
if 'resp_dir_override' in globals() and resp_dir_override:
resp_dir_candidates.append(resp_dir_override)
if len(resp_dir_candidates) == 0:
resp_dir_candidates.append(resp_dname)
resp_dirs = []
for d in resp_dir_candidates:
d_clean = d if d.endswith('/') else d + '/'
if os.path.isdir(d_clean):
resp_dirs.append(d_clean)
if len(resp_dirs) == 0:
raise FileNotFoundError('No valid response directory found in resp_dir_override / resp_dir_override_2 / resp_dname')
# keep compatibility with get_resp4ul_tab for first dir
calc_bat_ul.resp_dname = resp_dirs[0]
# collect .rsp from all dirs
all_rsp = []
for d in resp_dirs:
for f in os.listdir(d):
if f.endswith('.rsp'):
all_rsp.append((d, f))
if len(all_rsp) == 0:
raise ValueError(f'No .rsp files found in: {resp_dirs}')
# deduplicate by filename (first occurrence wins)
seen = set()
rsp_rows = []
for d, f in all_rsp:
if f in seen:
continue
seen.add(f)
rsp_rows.append((d, f))
rsp_dir_for_index = np.array([r[0] for r in rsp_rows])
fnames = np.array([r[1] for r in rsp_rows])
theta_values = np.array([float(f.split('_')[3]) for f in fnames])
phi_values = np.array([float(f.split('_')[5]) for f in fnames])
theta_str = np.array([f.split('_')[3] for f in fnames])
phi_str = np.array([f.split('_')[5] for f in fnames])
ras, decs = convert_theta_phi2radec(theta_values, phi_values, att_q)
def get_resp_tab_local(theta_s, phi_s, rsp_dir):
rsp_path = os.path.join(rsp_dir, f'NITRATES_alldet_theta_{theta_s}_phi_{phi_s}_.rsp')
if not os.path.exists(rsp_path):
raise FileNotFoundError(f'Response file not found: {rsp_path}')
return Table.read(rsp_path)
print(f'trigger_time_MET = {trigger_time:.3f}')
print(f'active detectors = {ndets_active}')
print(f'response directories = {resp_dirs}')
print(f'response points = {len(ras)}')
782 775
trigger_time_MET = 785996578.374
active detectors = 15537
response directories = ['/Users/sjs8171/Desktop/NITRATES/tests/nitrates_resp_dir/rsps4limits2/']
response points = 929
# ---- Convert rate UL -> flux UL at each response direction ----
rate_ul = compute_rate_upper_limit(bkg_obj, trigger_time, temporal_bin_s, sigma=sigma_ul)
print(f'rate upper limit ({sigma_ul:.1f} sigma, {temporal_bin_s:.3f} s) = {rate_ul:.4f} counts/s')
flux_ul_dirs = np.zeros(len(ras), dtype=float)
for i in range(len(ras)):
drm_tab = get_resp_tab_local(theta_str[i], phi_str[i], rsp_dir_for_index[i])
drm_matrix = drm_tab['MATRIX'][:, 0:4] * Ndet_ratio # 15-350 keV channels
if spectrum_model.lower() == 'band':
flux_ul = rate2band_eflux(
rate_ul,
drm_matrix,
drm_tab['ENERG_LO'],
drm_tab['ENERG_HI'],
band_alpha,
band_beta,
band_epeak_keV,
flux_elo_keV,
flux_ehi_keV,
)
elif spectrum_model.lower() == 'comp':
flux_ul = rate2comp_eflux(
rate_ul,
drm_matrix,
drm_tab['ENERG_LO'],
drm_tab['ENERG_HI'],
comp_alpha,
comp_epeak_keV,
flux_elo_keV,
flux_ehi_keV,
)
else:
raise ValueError("spectrum_model must be 'band' or 'comp'")
flux_ul_dirs[i] = flux_ul
print(f'min/max directional flux UL = {np.nanmin(flux_ul_dirs):.3e} / {np.nanmax(flux_ul_dirs):.3e} erg cm^-2 s^-1')
rate upper limit (5.0 sigma, 16.000 s) = 114.7970 counts/s
WARNING: hdu= was not specified but multiple tables are present, reading in first available table (hdu=1) [astropy.io.fits.connect]
min/max directional flux UL = 1.552e-08 / 3.120e-07 erg cm^-2 s^-1
# ---- Interpolate on HEALPix and write FITS ----
interp_ul = build_interpolator(ras, decs, flux_ul_dirs)
npix = hp.nside2npix(nside_out)
map_values = np.full(npix, np.nan, dtype=np.float64)
for pix in range(npix):
theta, phi = hp.pix2ang(nside_out, pix)
dec = np.degrees(np.pi / 2.0 - theta)
ra = np.degrees(phi)
map_values[pix] = interp_ul(ra, dec)
hp.write_map(output_fits, map_values, coord='C', overwrite=True, dtype=np.float64)
print(f'Wrote FITS sky map: {output_fits}')
Wrote FITS sky map: /Users/sjs8171/Desktop/acme_tutorials/nitrates/785996578_c0/ul_skymap_minimal.fits
# ! pip install ligo.skymap
from matplotlib.colors import LogNorm
# ---- Plot UL map (linear map, log colorbar) ----
import matplotlib.pyplot as plt
hpx, header = hp.read_map(output_fits, h=True)
valid = np.isfinite(hpx) & (hpx > 0)
if not np.any(valid):
raise ValueError('UL map has no positive finite pixels to plot in log scale.')
vmin = np.nanpercentile(hpx[valid], 1.0)
vmax = np.nanpercentile(hpx[valid], 99.5)
try:
import ligo.skymap.plot # noqa: F401
fig = plt.figure(figsize=(12, 7), dpi=100)
ax = plt.axes(projection='astro hours mollweide')
ax.grid()
image = ax.imshow_hpx(
(hpx, 'ICRS'),
cmap='viridis_r'
)
sm = plt.cm.ScalarMappable(cmap='viridis_r')
sm.set_norm(LogNorm(vmin=vmin, vmax=vmax))
sm.set_array([])
cbar = plt.colorbar(sm, ax=ax, orientation='horizontal', pad=0.05, shrink=0.8)
cbar.set_label('Flux upper limit (erg cm$^{-2}$ s$^{-1}$)')
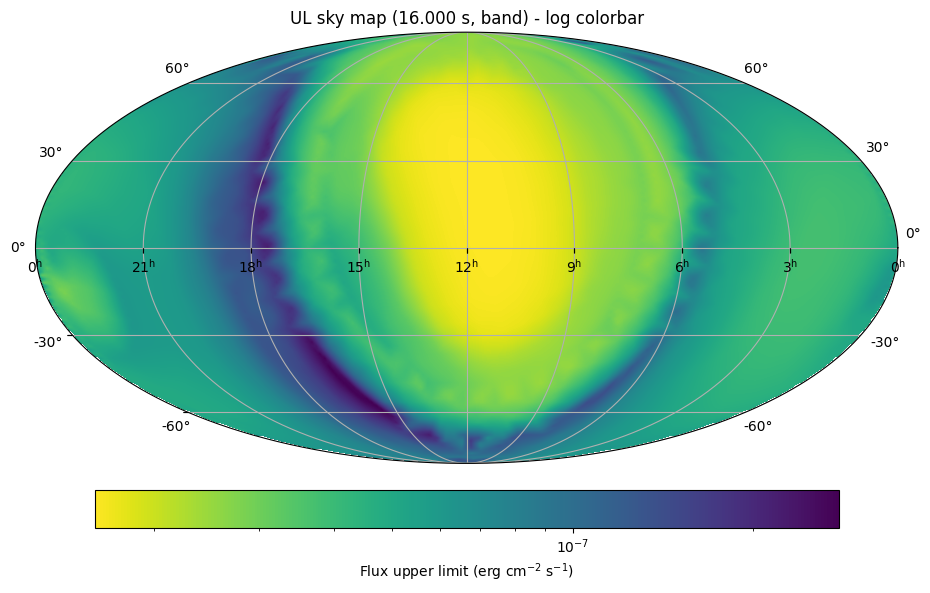
plt.title(f'UL sky map ({temporal_bin_s:.3f} s, {spectrum_model}) - log colorbar')
plt.show()
except ModuleNotFoundError:
hp.mollview(
hpx,
coord='C',
cmap='viridis_r',
norm='log',
min=vmin,
max=vmax,
title=f'UL sky map ({temporal_bin_s:.3f} s, {spectrum_model}) - log scale',
unit='Flux upper limit (erg cm$^{-2}$ s$^{-1}$)'
)
def ul_at_radec(ra_deg, dec_deg, ul_fits_path=output_fits, nest=False):
"""
Return flux upper limit at a given sky position (RA, Dec in degrees)
from the UL HEALPix FITS map.
"""
ul_map = hp.read_map(ul_fits_path, nest=nest, verbose=False)
nside = hp.get_nside(ul_map)
pix = hp.ang2pix(nside, ra_deg, dec_deg, lonlat=True, nest=nest)
return float(ul_map[pix])
# Example
ra_test, dec_test = 123.0, 50.0
ul_val = ul_at_radec(ra_test, dec_test)
print(f"UL at RA={ra_test:.3f}, Dec={dec_test:.3f} -> {ul_val:.3e} erg cm^-2 s^-1")
UL at RA=123.000, Dec=50.000 -> 6.202e-08 erg cm^-2 s^-1
/var/folders/5q/zf85wh2d0sxgdd55ws_1kk94pzxs0q/T/ipykernel_34482/4170089635.py:6: HealpyDeprecationWarning: "verbose" was deprecated in version 1.15.0 and will be removed in a future version.

