Sky-position scan with NITRATES#
This notebook performs a minimal non-imaging sky scan over a few trial positions and computes \(\sqrt{TS}\) at each position.
What this notebook does#
Load the bundled test dataset from
tests/nitrates_resp_dir(orNITRATES_RESP_DIRif already set).Build source + background likelihood objects.
Scan a small set of sky positions around a reference RA/Dec.
Profile over signal amplitude \(A\) at each position and compute:
Show a sky localization map
import os
import warnings
import io
import contextlib
# ── macOS kernel-crash prevention ────────────────────────────────────────────
# These MUST be set before ANY library that links OpenMP / MKL / Accelerate
# is imported. The root cause is that macOS loads multiple OpenMP runtimes
# simultaneously (libomp from conda-forge + libiomp5 from Intel/MKL), which
# corrupts Numba's JIT thread pool and causes a hard segfault.
#
# Key fix: switch Numba away from OpenMP entirely → use its own work-queue.
os.environ['NUMBA_THREADING_LAYER'] = 'workqueue' # ← the essential fix
os.environ['NUMBA_NUM_THREADS'] = '1'
os.environ['OMP_NUM_THREADS'] = '1'
os.environ['MKL_NUM_THREADS'] = '1'
os.environ['OPENBLAS_NUM_THREADS'] = '1'
os.environ['VECLIB_MAXIMUM_THREADS'] = '1' # macOS Accelerate
os.environ['KMP_DUPLICATE_LIB_OK'] = 'TRUE' # last-resort guard
# ─────────────────────────────────────────────────────────────────────────────
import logging
from pathlib import Path
import numpy as np
from astropy.io import fits
from nitrates.lib import (
get_conn,
get_info_tab,
mask_detxy,
convert_radec2thetaphi,
convert_theta_phi2radec,
)
from nitrates.response import RayTraces
from nitrates.models import (
Cutoff_Plaw_Flux,
Source_Model_InOutFoV,
CompoundModel,
Sig_Bkg_Model,
get_eflux_from_model,
)
from nitrates.llh_analysis import parse_bkg_csv, LLH_webins2, NLLH_ScipyMinimize_Wjacob
# Silence verbose internal output from NITRATES, astropy and root logger.
logging.basicConfig(level=logging.ERROR)
logging.getLogger().setLevel(logging.ERROR)
logging.getLogger('astropy').setLevel(logging.ERROR)
warnings.filterwarnings('ignore')
@contextlib.contextmanager
def suppress_output():
"""Redirect stdout/stderr to suppress print noise from NITRATES internals."""
with contextlib.redirect_stdout(io.StringIO()), contextlib.redirect_stderr(io.StringIO()):
yield
# Use the explicit local response directory
resp_dir = Path('/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir').resolve()
if not resp_dir.exists():
raise FileNotFoundError(f'NITRATES response directory not found: {resp_dir}')
os.environ['NITRATES_RESP_DIR'] = str(resp_dir)
rt_dir = resp_dir / 'ray_traces_detapp_npy'
solid_angle_dpi_path = resp_dir / 'solid_angle_dpi.npy'
print(f"Using responses from: {resp_dir}")
Using responses from: /Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir
trigid = '785996578_c0'
triggertime = '2025-11-28T04:22:21.610000'
# Define the literature/source sky position used throughout the tutorial.
source_ra_dec = (123, 50)
root = '/Users/samueleronchini/Desktop/nitrates_tutorial'
workdir = f'{root}/{trigid}'
# Trigger metadata
conn = get_conn(workdir + '/results.db')
info_tab = get_info_tab(conn)
trig = info_tab['trigtimeMET'][0]
# Event data and detector mask
ev_data = fits.open(workdir + '/filter_evdata.fits')[1].data
dmask = fits.open(workdir + '/detmask.fits')[0].data
mask_vals = mask_detxy(dmask, ev_data)
bl_dmask = dmask == 0.0
# Basic time/energy cuts
t_start = trig - 20
t_end = trig + 20
bl_ev = (
(ev_data['EVENT_FLAGS'] < 1)
& (ev_data['ENERGY'] <= 500.0)
& (ev_data['ENERGY'] >= 14.0)
& (mask_vals == 0.0)
& (ev_data['TIME'] <= t_end)
& (ev_data['TIME'] >= t_start)
)
filtered_ev_data = ev_data[bl_ev]
print(f"Selected {len(filtered_ev_data)} events over {t_end - t_start:.1f} s")
Selected 169204 events over 40.0 s
# Build the analysis energy bins in keV.
# `ebins0` contains the lower edge of each bin and `ebins1` the matching upper edge.
ebins0 = np.array([15.0, 24.0, 35.0, 48.0, 64.0])
ebins0 = np.append(ebins0, np.logspace(np.log10(84.0), np.log10(500.0), 5 + 1))[:-1]
ebins0 = np.round(ebins0, decimals=1)[:-1]
ebins1 = np.append(ebins0[1:], [350.0])
# Load the spacecraft attitude table and select the quaternion closest to the trigger time.
# This quaternion describes how the instrument was oriented on the sky at that moment.
with suppress_output():
attfile = fits.open(workdir + '/attitude.fits')[1].data
att_ind = np.argmin(np.abs(attfile['TIME'] - trig))
att_quat = attfile['QPARAM'][att_ind]
# Convert the source sky coordinates (RA, Dec) into instrument coordinates (theta, phi).
# The source response must be evaluated in the detector frame, not directly in RA/Dec.
theta, phi = convert_radec2thetaphi(*source_ra_dec, att_quat)
# Configure the spectral model parameters used as the initial source hypothesis.
# `A` is the normalization, `gamma` the photon index, and `Epeak` the cutoff/peak scale in keV.
flux_params = {'A': 1.0, 'gamma': 0.5, 'Epeak': 1e2}
flux_mod = Cutoff_Plaw_Flux(E0=100.0)
# Load the precomputed ray traces that encode detector shadowing across the field of view.
# Build the non-imaging source model using the spectral model, energy bins, detector mask,
# ray traces, and the response tables bundled with the tutorial dataset.
with suppress_output():
rt_obj = RayTraces(rt_dir)
sig_mod = Source_Model_InOutFoV(
flux_mod,
[ebins0, ebins1],
bl_dmask,
rt_obj,
use_deriv=True,
resp_tab_dname=resp_dir/'resp_tabs_ebins',
comp_flor_resp_dname=resp_dir/'comp_flor_resps',
hp_flor_resp_dname=resp_dir/'hp_flor_resps',
)
# Apply the instrument-frame sky position and spectral parameters to the source model.
sig_mod.set_theta_phi(theta, phi)
sig_mod.set_flux_params(flux_params)
# Print a compact summary so we can verify the selected attitude sample and derived geometry.
print(f"Pointing quaternion index: {att_ind}")
print(f"Theta/Phi (deg): {theta:.3f}, {phi:.3f}")
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(45.0, 54.0)
35.59031462549889
35.59031462549889
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_45.0_.fits
Pointing quaternion index: 175
Theta/Phi (deg): 52.335, 125.590
This cell prepares the source/background setup and chooses the analysis time bin from time_seeds.csv.
Loads attitude at trigger time and keeps the reference sky position.
Reads time_seeds.csv from the working directory.
Selects the best temporal seed (priority: snr, then TS, then \(\sqrt{TS}\)).
Converts the selected seed to absolute interval \([t_0, t_1]\).
Loads bkg_estimation.csv and extracts background parameters at the interval midpoint.
Builds reusable likelihood/model objects with the selected time window.
After this cell, all sky-position scans use the same best temporal bin from the seed file.
# --- Build source model ingredients and background model ---
# Attitude quaternion nearest trigger (needed for RA/Dec -> theta/phi conversion).
import glob
import pandas as pd
with suppress_output():
att = fits.open(resp_dir / 'attitude.fits')[1].data
att_idx = int(np.argmin(np.abs(att['TIME'] - trig)))
att_quat = att['QPARAM'][att_idx]
# Reference source position (can be changed if desired).
ra_ref, dec_ref = source_ra_dec[0], source_ra_dec[1]
# Choose analysis time bin from time_seeds.csv (best seed).
time_seeds_path = Path(workdir) / 'time_seeds.csv'
if not time_seeds_path.exists():
raise FileNotFoundError(f'time_seeds.csv not found: {time_seeds_path}')
time_seeds_df = pd.read_csv(time_seeds_path)
dur_col = 'duration' if 'duration' in time_seeds_df.columns else ('dur' if 'dur' in time_seeds_df.columns else None)
if dur_col is None:
raise KeyError("time_seeds.csv must contain 'duration' (or 'dur')")
# Prefer full-rates ranking if present.
if 'snr' in time_seeds_df.columns:
best_seed_idx = time_seeds_df['snr'].astype(float).idxmax()
elif 'TS' in time_seeds_df.columns:
best_seed_idx = time_seeds_df['TS'].astype(float).idxmax()
elif 'sqrtTS' in time_seeds_df.columns:
best_seed_idx = time_seeds_df['sqrtTS'].astype(float).idxmax()
else:
raise KeyError("time_seeds.csv must contain one ranking column among: 'snr', 'TS', 'sqrtTS'")
best_seed = time_seeds_df.loc[best_seed_idx]
dur_best = float(best_seed[dur_col])
if 'time' in time_seeds_df.columns:
t0 = float(best_seed['time'])
elif 'dt' in time_seeds_df.columns:
t0 = float(trig) + float(best_seed['dt'])
else:
raise KeyError("time_seeds.csv must contain either 'time' or 'dt'")
t1 = t0 + dur_best
print(
f"Selected best time seed from {time_seeds_path.name}: "
f"t0-trig={t0 - float(trig):+.3f} s, dur={dur_best:.3f} s"
)
# Spectral shape: take gamma/Epeak from split-rates results (same or nearest time bin).
split_files = sorted(glob.glob(str(Path(workdir) / 'rates_llh_res_*.csv')))
if not split_files:
raise FileNotFoundError('No split-rate files found (rates_llh_res_*.csv). Run split-rates first.')
split_df = pd.concat([pd.read_csv(f) for f in split_files], ignore_index=True)
if 'gamma' not in split_df.columns or 'Epeak' not in split_df.columns:
raise KeyError("Split-rate table must contain 'gamma' and 'Epeak' columns")
# Match split-rates row to selected time seed.
cand = split_df.copy()
# Prefer exact match by timeID if available in both tables.
if 'timeID' in cand.columns and 'timeID' in best_seed.index:
cand = cand[cand['timeID'].astype(float) == float(best_seed['timeID'])]
# If timeID match is empty/not available, use nearest in (time, duration).
if len(cand) == 0:
if 'time' in split_df.columns and 'dur' in split_df.columns:
tmp = split_df.copy()
tmp['_metric'] = (tmp['time'].astype(float) - float(t0)).abs() + (tmp['dur'].astype(float) - float(dur_best)).abs()
min_metric = float(tmp['_metric'].min())
cand = tmp[tmp['_metric'] == min_metric].copy()
else:
cand = split_df.copy()
# Choose highest-significance split-rate row among candidates.
rank_col = 'TS' if 'TS' in cand.columns else ('sqrtTS' if 'sqrtTS' in cand.columns else None)
if rank_col is not None:
best_split = cand.loc[cand[rank_col].astype(float).idxmax()]
else:
best_split = cand.iloc[0]
gamma_best = float(best_split['gamma'])
epeak_best = float(best_split['Epeak'])
flux_mod = Cutoff_Plaw_Flux(E0=100.0)
flux_shape = {'A': 1.0, 'gamma': gamma_best, 'Epeak': epeak_best}
print(f"Using split-rates spectral shape: gamma={gamma_best:.3f}, Epeak={epeak_best:.2f} keV")
# Background model and parameters at analysis time midpoint.
bkg_csv_path = Path(workdir) / 'bkg_estimation.csv'
with suppress_output():
bkg_df, bkg_name, _, bkg_mod, ps_mods = parse_bkg_csv(
bkg_csv_path,
np.load(solid_angle_dpi_path),
ebins0,
ebins1,
bl_dmask,
rt_dir,
)
tmid = 0.5 * (t0 + t1)
bkg_row = bkg_df.iloc[np.argmin(np.abs(bkg_df['time'] - tmid))]
# Some background CSV files store columns with a prefix (e.g. "Background_bkg_rate_0")
# while model parameter names may be unprefixed (e.g. "bkg_rate_0").
def _get_bkg_value(row, pname, prefix):
if pname in row.index:
return row[pname]
pref_name = f"{prefix}{pname}" if prefix else pname
if pref_name in row.index:
return row[pref_name]
if pname.startswith('Background_'):
no_pref = pname[len('Background_'):]
if no_pref in row.index:
return row[no_pref]
raise KeyError(f"Background parameter '{pname}' not found in CSV columns")
bkg_params = {p: _get_bkg_value(bkg_row, p, bkg_name) for p in bkg_mod.param_names}
# Reusable objects that do not depend on trial sky position.
with suppress_output():
rt_obj = RayTraces(rt_dir)
llh = LLH_webins2(filtered_ev_data, ebins0, ebins1, bl_dmask, has_err=True)
llh.set_time(t0, t1)
print('Source/background setup complete.')
Selected best time seed from time_seeds.csv: t0-trig=-1.024 s, dur=4.096 s
Using split-rates spectral shape: gamma=-0.100, Epeak=411.33 keV
Source/background setup complete.
eval_sky_position#
This helper evaluates one trial sky coordinate and returns the detection significance for that point.
Receives trial \((\mathrm{RA},\mathrm{Dec})\) and clips/wraps values to valid ranges.
Converts sky coordinates to instrument coordinates \((\theta,\phi)\) using the trigger-time attitude.
Builds a trial source model at that sky position with fixed spectral shape.
Combines source + background models and attaches them to the likelihood object.
Computes background-only NLLH with \(A\approx 0\).
Profiles the likelihood over amplitude
A(all other parameters fixed).Computes \(\sqrt{TS}=\sqrt{2(\mathrm{NLLH}_{\mathrm{bkg}}-\mathrm{NLLH}_{\mathrm{best}})}\) and returns a result dictionary.
Returned fields include position, best-fit amplitude, NLLH values, and sqrtTS.
# --- Helper: evaluate one sky position and return sqrtTS ---
def eval_sky_position(ra_deg: float, dec_deg: float):
"""
Evaluate one trial (RA, Dec):
1) Convert to instrument coordinates (theta, phi).
2) Build source+background model with fixed spectral shape.
3) Profile likelihood over amplitude A only.
4) Compute sqrtTS relative to background-only hypothesis.
"""
# Keep values in physical ranges for robust scanning.
ra_deg = float(ra_deg % 360.0)
dec_deg = float(np.clip(dec_deg, -89.0, 89.0))
# Sky -> instrument frame.
theta, phi = convert_radec2thetaphi(ra_deg, dec_deg, att_quat)
# Build trial source model at this sky location.
with suppress_output():
sig_mod = Source_Model_InOutFoV(
flux_mod,
[ebins0, ebins1],
bl_dmask,
rt_obj,
use_deriv=True,
resp_tab_dname=resp_dir / 'resp_tabs_ebins',
comp_flor_resp_dname=resp_dir / 'comp_flor_resps',
hp_flor_resp_dname=resp_dir / 'hp_flor_resps',
)
sig_mod.set_theta_phi(float(theta), float(phi))
sig_mod.set_flux_params(flux_shape)
# Combine source + background and set parameters.
sig_bkg = Sig_Bkg_Model(bl_dmask, sig_mod, bkg_mod, use_deriv=True)
sig_bkg.set_bkg_params(bkg_params)
sig_bkg.set_sig_params({
'A': 1.0,
'gamma': float(flux_shape['gamma']),
'Epeak': float(flux_shape['Epeak']),
'theta': float(theta),
'phi': float(phi),
})
# Attach model to LLH object.
llh.set_model(sig_bkg)
# Background-only NLLH (A ~ 0).
nllh_bkg = -float(llh.get_logprob({'A': 1e-10}))
# Profile over A with all other parameters fixed.
miner = NLLH_ScipyMinimize_Wjacob('')
miner.set_llh(llh)
miner.set_fixed_params(miner.param_names, fixed=True)
miner.set_fixed_params(['A'], fixed=False)
bf_vals, nllh_vals, _ = miner.minimize()
A_best = float(np.atleast_1d(bf_vals[0])[0])
nllh_best = float(nllh_vals[0])
# sqrtTS from profile likelihood ratio.
sqrt_ts = float(np.sqrt(max(2.0 * (nllh_bkg - nllh_best), 0.0)))
return {
'ra': ra_deg,
'dec': dec_deg,
'theta': float(theta),
'phi': float(phi),
'A_best': A_best,
'nllh_bkg': nllh_bkg,
'nllh_best': nllh_best,
'sqrtTS': sqrt_ts,
}
import pandas as pd
# --- Define a 3×3 grid of trial sky positions around the reference point ---
offsets = np.linspace(-35, 35, 4)
trial_positions = [
(ra_ref + dra, dec_ref + ddec)
for ddec in offsets
for dra in offsets
]
rows = []
for ra_i, dec_i in trial_positions:
try:
rows.append(eval_sky_position(ra_i, dec_i))
except Exception as e:
# If one position is unsupported by response coverage, continue scanning others.
rows.append({
'ra': float(ra_i),
'dec': float(dec_i),
'theta': np.nan,
'phi': np.nan,
'A_best': np.nan,
'nllh_bkg': np.nan,
'nllh_best': np.nan,
'sqrtTS': np.nan,
'error': str(e),
})
scan_df = pd.DataFrame(rows)
scan_df = scan_df.sort_values('sqrtTS', ascending=False, na_position='last').reset_index(drop=True)
display_cols = ['ra', 'dec', 'A_best', 'sqrtTS', 'nllh_bkg', 'nllh_best']
print('Sky scan results (sorted by sqrtTS):')
display(scan_df[display_cols])
if scan_df['sqrtTS'].notna().any():
best = scan_df.loc[scan_df['sqrtTS'].idxmax()]
print(
f"Best trial position: RA={best['ra']:.3f}, Dec={best['dec']:.3f}, "
f"sqrtTS={best['sqrtTS']:.3f}, A_best={best['A_best']:.3e}"
)
else:
print('No valid sky points were evaluated.')
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(63.0, 72.0)
33.717838404782015
11.282161595217985
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_72.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_72.0_phi_15.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(36.0, 45.0)
21.476676602570535
23.523323397429465
max rt: 11579.5469
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(18.0, 27.0)
0.23358637578198227
44.76641362421802
max rt: 9482.4775
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(9.0, 18.0)
27.57411664722798
17.42588335277202
max rt: 3489.5300
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_9.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_9.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
13.88001070710095
13.88001070710095
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(36.0, 45.0)
10.766206533317359
10.766206533317359
max rt: 74922.2578
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_15.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(18.0, 27.0)
16.29342971053245
16.29342971053245
max rt: 39724.0039
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_18.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(3.0, 9.0)
33.402780231965664
11.597219768034336
max rt: 13883.1631
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_3.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_3.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_9.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_9.0_phi_15.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
42.069756714750156
42.069756714750156
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(36.0, 45.0)
0.5845092272479633
44.41549077275204
max rt: 105068.5781
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_45.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
11.68553895743753
33.31446104256247
max rt: 69391.7734
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
30.660793021071754
14.339206978928246
max rt: 51302.4922
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_15.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
24.611820211689405
20.388179788310595
max rt: 203659.6250
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
25.796727099852205
19.203272900147795
max rt: 179698.3281
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
27.754533602557387
17.245466397442613
max rt: 161413.0469
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_30.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(54.0, 63.0)
30.14959002791278
14.85040997208722
max rt: 149286.1562
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_54.0_phi_15.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_0.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_63.0_phi_15.0_.fits
Sky scan results (sorted by sqrtTS):
| ra | dec | A_best | sqrtTS | nllh_bkg | nllh_best | |
|---|---|---|---|---|---|---|
| 0 | 88.000000 | 15.000000 | 0.010049 | 16.570776 | 51767.325646 | 51630.030332 |
| 1 | 158.000000 | 38.333333 | 0.006979 | 16.276956 | 51767.325645 | 51634.855998 |
| 2 | 111.333333 | 15.000000 | 0.006873 | 16.210564 | 51767.325645 | 51635.934451 |
| 3 | 88.000000 | 85.000000 | 0.006788 | 15.747367 | 51767.325645 | 51643.335867 |
| 4 | 111.333333 | 85.000000 | 0.006873 | 15.540036 | 51767.325645 | 51646.579283 |
| 5 | 134.666667 | 61.666667 | 0.008508 | 15.536782 | 51767.325646 | 51646.629841 |
| 6 | 158.000000 | 61.666667 | 0.007042 | 15.471196 | 51767.325645 | 51647.646686 |
| 7 | 158.000000 | 85.000000 | 0.006715 | 15.458336 | 51767.325645 | 51647.845566 |
| 8 | 158.000000 | 15.000000 | 0.005905 | 15.428516 | 51767.325645 | 51648.306096 |
| 9 | 134.666667 | 85.000000 | 0.006800 | 15.363699 | 51767.325645 | 51649.304018 |
| 10 | 111.333333 | 38.333333 | 0.008080 | 15.060840 | 51767.325646 | 51653.911190 |
| 11 | 111.333333 | 61.666667 | 0.008689 | 14.992566 | 51767.325646 | 51654.937130 |
| 12 | 134.666667 | 15.000000 | 0.006503 | 14.987763 | 51767.325645 | 51655.009123 |
| 13 | 134.666667 | 38.333333 | 0.006435 | 14.612812 | 51767.325645 | 51660.558501 |
| 14 | 88.000000 | 38.333333 | 0.007430 | 13.528740 | 51767.325646 | 51675.812237 |
| 15 | 88.000000 | 61.666667 | 0.006187 | 12.777149 | 51767.325646 | 51685.697876 |
Best trial position: RA=88.000, Dec=15.000, sqrtTS=16.571, A_best=1.005e-02
# --- 2D map: sqrtTS on the RA/Dec grid ---
import matplotlib.pyplot as plt
import numpy as np
valid = scan_df['sqrtTS'].notna()
if valid.any():
df_v = scan_df.loc[valid].copy()
# Reconstruct the 2D grid (RA increases left → invert x-axis later)
ra_vals = np.sort(df_v['ra'].unique())
dec_vals = np.sort(df_v['dec'].unique())
grid = np.full((len(dec_vals), len(ra_vals)), np.nan)
ra_idx = {v: i for i, v in enumerate(ra_vals)}
dec_idx = {v: i for i, v in enumerate(dec_vals)}
for _, row in df_v.iterrows():
grid[dec_idx[row['dec']], ra_idx[row['ra']]] = row['sqrtTS']
fig, ax = plt.subplots(figsize=(6.5, 5.2))
im = ax.pcolormesh(
ra_vals, dec_vals, grid,
cmap='viridis', shading='nearest',
)
plt.colorbar(im, ax=ax, label=r'$\sqrt{TS}$')
# Mark best position
best = scan_df.loc[scan_df['sqrtTS'].idxmax()]
ax.scatter(
best['ra'], best['dec'],
marker='*', s=280, color='tab:red',
edgecolors='white', linewidths=0.8,
zorder=5, label=f"Best ({best['sqrtTS']:.2f})",
)
# Mark reference position
ax.scatter(
[ra_ref], [dec_ref],
marker='+', s=160, color='white',
linewidths=1.5, zorder=5, label='Reference',
)
ax.set_xlabel('RA [deg]')
ax.set_ylabel('Dec [deg]')
ax.set_title(r'Sky scan — $\sqrt{TS}$ map (3×3 grid)')
ax.invert_xaxis()
ax.legend(loc='upper right', fontsize=9)
plt.tight_layout()
plt.show()
else:
print('No valid points to plot.')
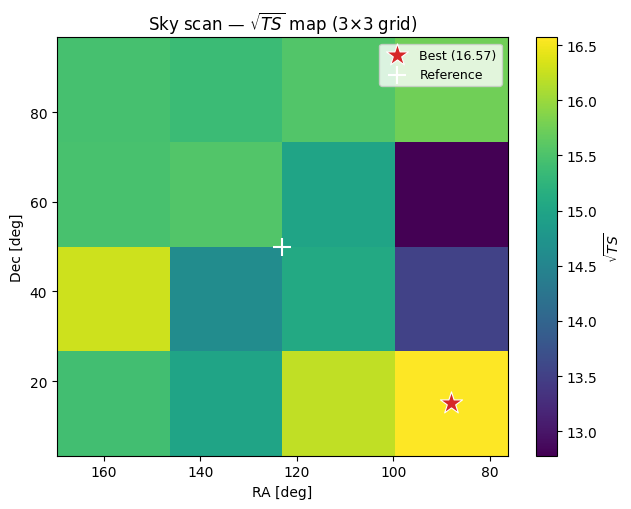
Notes#
This is intentionally minimal: it scans only 5 trial positions.
Spectral shape (
gamma,Epeak) is fixed; only amplitude \(A\) is profiled.To increase localization detail, enlarge the trial grid (more RA/Dec offsets).
If some trial points fail, they are reported as NaN (typically response-coverage limits).
Extra section: max TS on a joint sky + spectral grid#
In this final section we do a compact joint grid scan over:
sky positions (RA/Dec around the reference), and
spectral shape parameters (
gamma,Epeak).
For each grid point we:
Build the source model at that sky position.
Fix (
gamma,Epeak) to trial values.Profile only over amplitude
A.Compute
TS = 2(\mathrm{NLLH}_{bkg} - \mathrm{NLLH}_{sig+bkg})andsqrtTS.
This is slower than the essential scan, so the default grids below are intentionally small.
# --- Joint sky + spectral grid scan (profile over A) ---
# Small default grids (edit these to trade speed vs detail)
ra_offsets_deg = np.array([-1.0, 0.0, 1.0])
dec_offsets_deg = np.array([-1.0, 0.0, 1.0])
gamma_grid = np.array([0.0, 0.5, 1.0])
epeak_grid = np.array([70.0, 100.0, 150.0])
# Reuse event selection and time window from previous sections
llh_joint = LLH_webins2(filtered_ev_data, ebins0, ebins1, bl_dmask, has_err=True)
llh_joint.set_time(t0, t1)
joint_rows = []
for dra in ra_offsets_deg:
for ddec in dec_offsets_deg:
ra_i = float((ra_ref + dra) % 360.0)
dec_i = float(np.clip(dec_ref + ddec, -89.0, 89.0))
try:
theta_i, phi_i = convert_radec2thetaphi(ra_i, dec_i, att_quat)
# Build source model once per sky position
with suppress_output():
sig_mod_i = Source_Model_InOutFoV(
flux_mod,
[ebins0, ebins1],
bl_dmask,
rt_obj,
use_deriv=True,
resp_tab_dname=resp_dir / 'resp_tabs_ebins',
comp_flor_resp_dname=resp_dir / 'comp_flor_resps',
hp_flor_resp_dname=resp_dir / 'hp_flor_resps',
)
sig_mod_i.set_theta_phi(float(theta_i), float(phi_i))
for g_i in gamma_grid:
for ep_i in epeak_grid:
try:
# Fixed spectral shape for this trial point
trial_flux = {'A': 1.0, 'gamma': float(g_i), 'Epeak': float(ep_i)}
sig_mod_i.set_flux_params(trial_flux)
sig_bkg_i = Sig_Bkg_Model(bl_dmask, sig_mod_i, bkg_mod, use_deriv=True)
sig_bkg_i.set_bkg_params(bkg_params)
sig_bkg_i.set_sig_params({
'A': 1.0,
'gamma': float(g_i),
'Epeak': float(ep_i),
'theta': float(theta_i),
'phi': float(phi_i),
})
llh_joint.set_model(sig_bkg_i)
# Background-only reference at this trial configuration
nllh_bkg_i = -float(llh_joint.get_logprob({'A': 1e-10}))
# Profile over A only
miner_i = NLLH_ScipyMinimize_Wjacob('')
miner_i.set_llh(llh_joint)
miner_i.set_fixed_params(miner_i.param_names, fixed=True)
miner_i.set_fixed_params(['A'], fixed=False)
bf_vals_i, nllh_vals_i, _ = miner_i.minimize()
A_best_i = float(np.atleast_1d(bf_vals_i[0])[0])
nllh_best_i = float(nllh_vals_i[0])
ts_i = float(max(2.0 * (nllh_bkg_i - nllh_best_i), 0.0))
sqrt_ts_i = float(np.sqrt(ts_i))
joint_rows.append({
'ra': ra_i,
'dec': dec_i,
'theta': float(theta_i),
'phi': float(phi_i),
'gamma': float(g_i),
'Epeak': float(ep_i),
'A_best': A_best_i,
'nllh_bkg': nllh_bkg_i,
'nllh_best': nllh_best_i,
'TS': ts_i,
'sqrtTS': sqrt_ts_i,
})
except Exception as e_inner:
joint_rows.append({
'ra': ra_i,
'dec': dec_i,
'theta': float(theta_i),
'phi': float(phi_i),
'gamma': float(g_i),
'Epeak': float(ep_i),
'A_best': np.nan,
'nllh_bkg': np.nan,
'nllh_best': np.nan,
'TS': np.nan,
'sqrtTS': np.nan,
'error': str(e_inner),
})
except Exception as e_sky:
# If this sky position is unsupported, register all spectral points as failed
for g_i in gamma_grid:
for ep_i in epeak_grid:
joint_rows.append({
'ra': ra_i,
'dec': dec_i,
'theta': np.nan,
'phi': np.nan,
'gamma': float(g_i),
'Epeak': float(ep_i),
'A_best': np.nan,
'nllh_bkg': np.nan,
'nllh_best': np.nan,
'TS': np.nan,
'sqrtTS': np.nan,
'error': str(e_sky),
})
joint_df = pd.DataFrame(joint_rows)
valid_joint = joint_df.dropna(subset=['TS']).reset_index(drop=True)
print(f'Total trial points: {len(joint_df)}')
print(f'Valid trial points: {len(valid_joint)}')
if len(valid_joint) == 0:
print('No valid joint-grid points were evaluated.')
else:
best_joint = valid_joint.loc[valid_joint['TS'].idxmax()]
print('\nBest point in joint sky+spectral grid:')
print(
f"RA={best_joint['ra']:.3f}, Dec={best_joint['dec']:.3f}, "
f"gamma={best_joint['gamma']:.3f}, Epeak={best_joint['Epeak']:.1f} keV"
)
print(
f"A_best={best_joint['A_best']:.3e}, TS={best_joint['TS']:.3f}, "
f"sqrtTS={best_joint['sqrtTS']:.3f}"
)
display_cols_joint = ['ra', 'dec', 'gamma', 'Epeak', 'A_best', 'TS', 'sqrtTS']
display(valid_joint.sort_values('TS', ascending=False).head(15)[display_cols_joint])
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
31.093602915084034
31.093602915084034
max rt: 51404.4023
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
32.76158022149434
32.76158022149434
max rt: 53960.8984
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
34.39382835789979
34.39382835789979
max rt: 56533.2461
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
31.575517335462337
31.575517335462337
max rt: 50247.6250
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
33.25478506588399
33.25478506588399
max rt: 52766.4023
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
34.89563373052482
34.89563373052482
max rt: 55302.5742
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
32.088416157450126
32.088416157450126
max rt: 49138.7461
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
33.77814263241669
33.77814263241669
max rt: 51620.2500
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
(0.0, 30.48, -14.117)
(0.0, 30.48, -14.117)
(27.0, 36.0)
35.42663622314768
35.42663622314768
max rt: 54120.6016
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_27.0_phi_45.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_30.0_.fits
initing ResponseDPI, with fname
/Users/samueleronchini/Desktop/NITRATES/tests/nitrates_resp_dir/resp_tabs_ebins/drm_theta_36.0_phi_45.0_.fits
Total trial points: 81
Valid trial points: 81
Best point in joint sky+spectral grid:
RA=122.000, Dec=49.000, gamma=0.000, Epeak=150.0 keV
A_best=4.103e-02, TS=217.962, sqrtTS=14.764
| ra | dec | gamma | Epeak | A_best | TS | sqrtTS | |
|---|---|---|---|---|---|---|---|
| 2 | 122.0 | 49.0 | 0.0 | 150.0 | 0.041032 | 217.961614 | 14.763523 |
| 11 | 122.0 | 50.0 | 0.0 | 150.0 | 0.039875 | 208.171044 | 14.428134 |
| 20 | 122.0 | 51.0 | 0.0 | 150.0 | 0.039340 | 205.484225 | 14.334721 |
| 5 | 122.0 | 49.0 | 0.5 | 150.0 | 0.025361 | 197.153532 | 14.041137 |
| 14 | 122.0 | 50.0 | 0.5 | 150.0 | 0.024465 | 186.771093 | 13.666422 |
| 23 | 122.0 | 51.0 | 0.5 | 150.0 | 0.024020 | 183.236223 | 13.536477 |
| 29 | 123.0 | 49.0 | 0.0 | 150.0 | 0.035498 | 181.115820 | 13.457928 |
| 38 | 123.0 | 50.0 | 0.0 | 150.0 | 0.035428 | 179.895443 | 13.412511 |
| 47 | 123.0 | 51.0 | 0.0 | 150.0 | 0.034915 | 175.941729 | 13.264303 |
| 65 | 124.0 | 50.0 | 0.0 | 150.0 | 0.033736 | 175.117663 | 13.233203 |
| 74 | 124.0 | 51.0 | 0.0 | 150.0 | 0.033610 | 173.415560 | 13.168734 |
| 56 | 124.0 | 49.0 | 0.0 | 150.0 | 0.033449 | 173.056582 | 13.155097 |
| 8 | 122.0 | 49.0 | 1.0 | 150.0 | 0.014568 | 168.775743 | 12.991372 |
| 32 | 123.0 | 49.0 | 0.5 | 150.0 | 0.021525 | 161.151159 | 12.694533 |
| 41 | 123.0 | 50.0 | 0.5 | 150.0 | 0.021406 | 159.755010 | 12.639423 |

